What are protein interaction networks?
Protein interaction networks are diagrams that visually present the interactions with other proteins that a protein of interest has in cells. A protein interaction depicted in these networks could indicate that the proteins bind to one another, affect the functionality of one another, or that they are both part of a signaling cascade [1]. Protein interaction networks can be constructed experimentally (through the use of BioID, SILAC or iTRAQ), or computationally by using STRING. STRING is able to construct interaction networks by showing protein interactions that are already determined experimentally or that are predicted by analyzing protein domains and similar existing interactions [2]. The information that can be gathered from a protein interaction network can be used to find patterns in the proteins that the protein of interest interacts with and these can be sorted by relevant GO terms. This will help determine the role and function of the protein of interest.
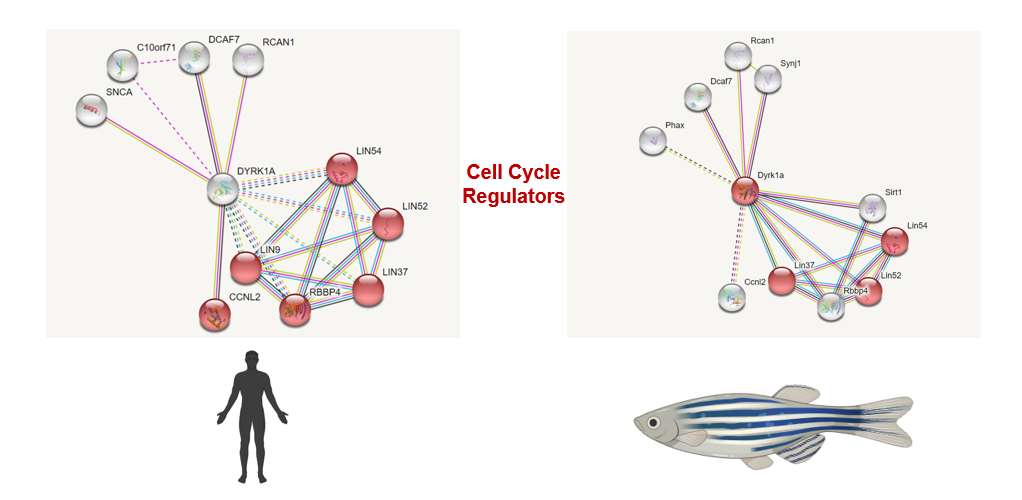
DYRK1A interaction networks
Discussion
The interactions between DYRK1A and transcription factors implicated in neuronal differentiation and development are well studied [3]. It has been observed that DYRK1A also is associated with reduced neural cell proliferation, however this particular association is not well-characterized. When analyzing the STRING findings shown above, I found that DYRK1A interacts with a number of cell cycle regulatory proteins. This especially interested me because interacting with cell cycle proteins could lead to a possible mechanism in which DYRK1A is able to modulate neural cell proliferation. Another important finding was that the interactions between DYRK1A and important cell cycle regulators in both humans and zebrafish are very similar. This further proves that zebrafish would be an ideal model organism to conduct my studies of DYRK1A and its role in neural cell proliferation. Based on these findings, I would like to confirm these DYRK1A protein interactions experimentally by the use of BioID, and this could lead to a better understanding of how DYRK1A reduced neural cell proliferation.
References
[1] Protein-protein interaction networks. (n.d.) EMBL-EBI. Retrieved from: https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
[2] STRING. https://string-db.org/cgi/network.pl?taskId=FMN45rVBeYzb
[3] Liu, Y., Lin, Z., Liu, M., Wang, H., & Sun, H. (2017). Overexpression of DYRK1A, a Down Syndrome Candidate gene, Impairs Primordial Germ Cells Maintenance and Migration in zebrafish. Scientific reports, 7(1), 15313. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5681638/
[2] STRING. https://string-db.org/cgi/network.pl?taskId=FMN45rVBeYzb
[3] Liu, Y., Lin, Z., Liu, M., Wang, H., & Sun, H. (2017). Overexpression of DYRK1A, a Down Syndrome Candidate gene, Impairs Primordial Germ Cells Maintenance and Migration in zebrafish. Scientific reports, 7(1), 15313. Retrieved from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5681638/
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.