What is Transcriptomics?
Transcriptomics is the analysis of the transcriptome. The transcriptome refers to all the sets of RNA transcripts within a single or population of cells [1]. Transcriptomics can be used to study RNA molecules within a sample of tissue under certain experimental conditions. Transciptomics studies that are conducted under differing experimental conditions can elucidate what genes are expressed and where they are expressed to better understand which gene expression profiles are correlated with the observed phenotypes [2]. Additionally, the function of particular genes and less-known gene expression patterns can be determined through the use of transcriptomics.
There are multiple techniques that can be used to analyze transcriptomes such as microarrays, RNA sequencing, and single-cell RNA sequencing. Microarrays are used to see the relative levels of expression among a set of genes between different experimental conditions [3]. RNA sequencing is an extremely useful technique because it can be used to analyze the transcriptome of whole tissues [3]. Single-cell RNA sequencing is a more recently developed technique that allows for a more specific analysis of the transcriptome between different time points and different experimental conditions [3].
There are multiple techniques that can be used to analyze transcriptomes such as microarrays, RNA sequencing, and single-cell RNA sequencing. Microarrays are used to see the relative levels of expression among a set of genes between different experimental conditions [3]. RNA sequencing is an extremely useful technique because it can be used to analyze the transcriptome of whole tissues [3]. Single-cell RNA sequencing is a more recently developed technique that allows for a more specific analysis of the transcriptome between different time points and different experimental conditions [3].
DYRK1A Expression Patterns
Discussion
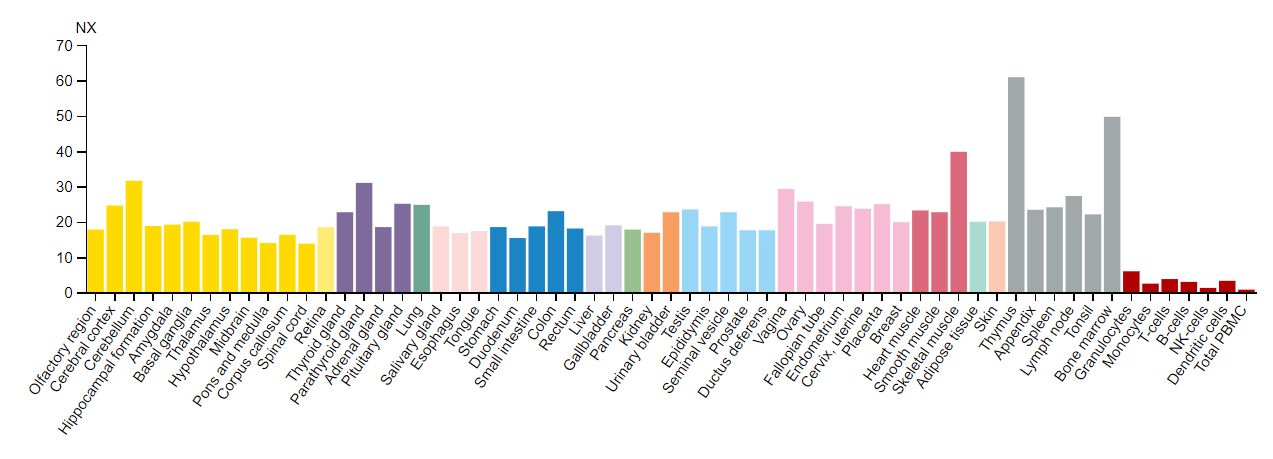
As seen in the tissue expression profile above, DYRK1A is highly expressed in several tissue types. This makes sense because DYRK1A has a wide variety of functions and has a wide range of targets within cells. It is especially interesting that DYRK1A is highly expressed in the thymus and bone marrow - tissue types that associated with the generation of white blood cells. Interestingly, DYRK1A has a well-characterized role in the development of lymphocytes, hence the high expression levels in the thymus and bone marrow. In fact, DYRK1A has well-characterized and unique roles in several different tissue types. However, I will focus on DYRK1A expression within the central nervous system to see its role in neurodegenerative disorders. DYRK1A is highly expressed in the central nervous system, especially within the cerebral cortex and the cerebellum. The cerebral cortex is responsible for higher level processing such as speech and decision making while the cerebellum is responsible for coordination and muscle activity. DYRK1A and its inhibitory nature of transcription factors in these tissue regions can explain the reduced higher level processing and decreased muscle tone seen in individuals with Down syndrome.
References
1. https://www.sciencedirect.com/topics/neuroscience/transcriptomics
2. https://www.proteinatlas.org/ENSG00000157540-DYRK1A/summary/rna
3. NIH. (August 2015). Transcriptome. Retrieved from: https://www.genome.gov/13014330/transcriptome-fact-sheet/
2. https://www.proteinatlas.org/ENSG00000157540-DYRK1A/summary/rna
3. NIH. (August 2015). Transcriptome. Retrieved from: https://www.genome.gov/13014330/transcriptome-fact-sheet/
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.